CLOCTools.github.io
CLOCTools
Welcome! CLOCTools is a library of tools for closed-loop neuroscience developed by Christopher Rozell’s and Garrett Stanley’s labs at Georgia Tech. CLOCTools includes core algorithms for state-space and state-switching control as well as tools for implementation in real-time experiments. Additionally, you’ll find a list of relevant publications and related tools for simulation, modeling, and analysis.
Overview
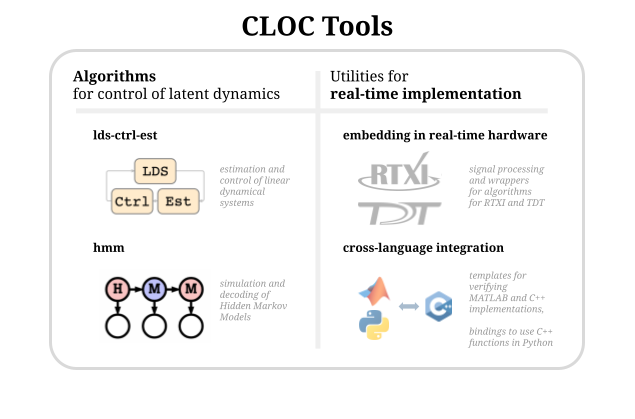
CLOCTools algorithms include linear dynamical systems control estimation (ldsCtrlEst) and hidden Markov model switching linear dynamical systems (hmm). For implementation, CLOC tools has a variety of software for interfacing with the the real time tools RTXI and TDT along with tools for interfacing across programming languages.
Core Algorithms
State-space dynamics (GLDS, PLDS) CLOCTools/ldsCtrlEst
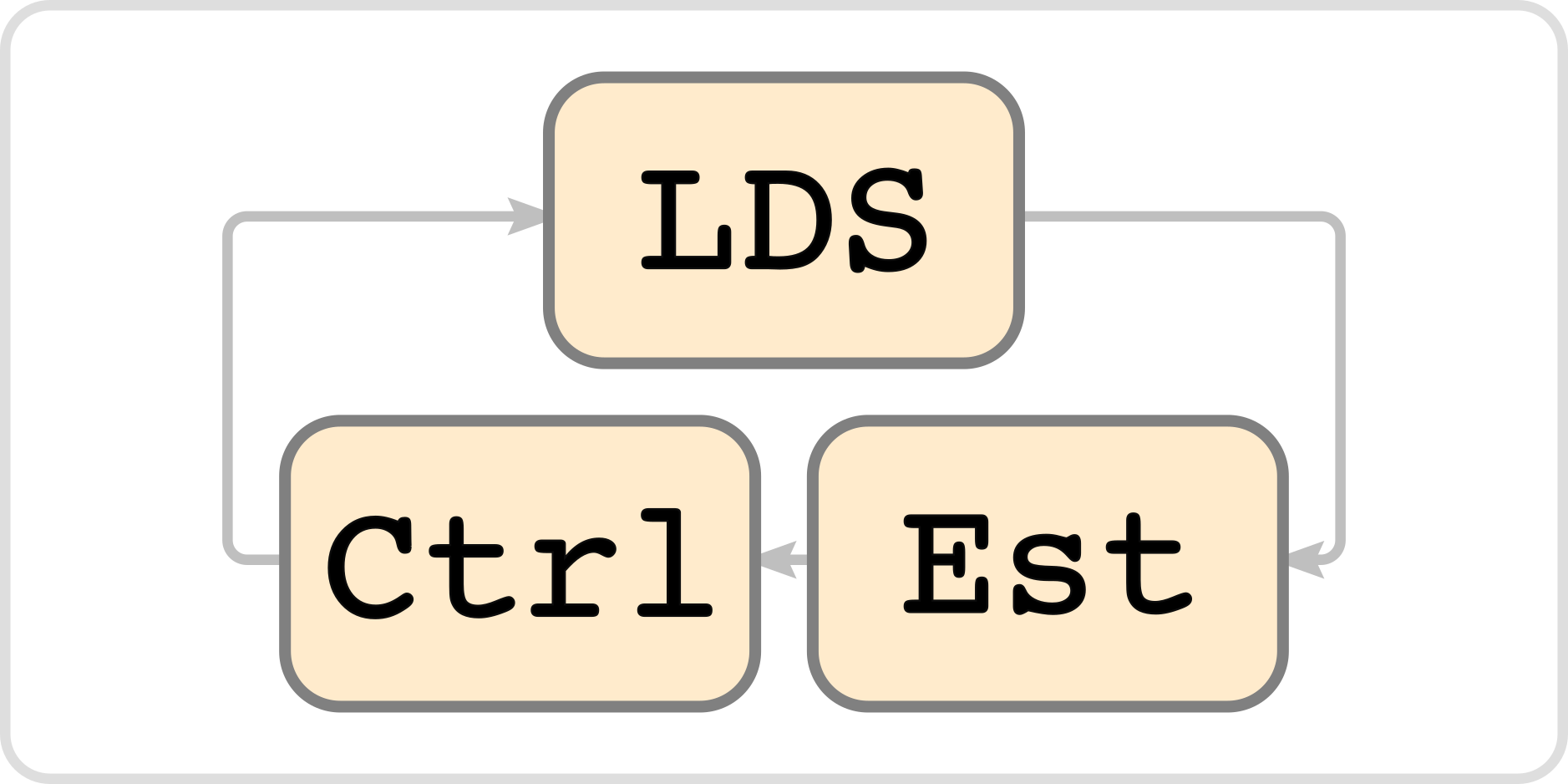
ldsCtrlEst is a C++ library for estimation and control of linear dynamical systems (LDS) with Gaussian or Poisson observations. It is meant to provide the functionality necessary to implement feedback control of linear dynamical systems experimentally. This library was originally developed for the task of controlling neuronal activity using spike count data as feedback and optogenetic inputs for control. ldsCtrlEst has both estimation and feedback control with parameter adaptive estimation, switched estimate and control, and can use both Gaussian and Poisson observation models.
The repository also provides an alternate MATLAB implementation, MATLAB bindings to model fitting functions, and Python bindings to the entire library.
example scripts:
- GLDS control -
eg_glds_ctrl(.cpp|.py) - tutorial - GLDS control of PLDS -
eg_glds_du_plds_ctrl(.cpp|.py)- change in control (du) is being updated, rather than amplitude (u).
- PLDS control -
eg_plds_ctrl(.cpp|.py) - PLDS estimation -
eg_plds_est(.cpp|.py) - tutorial - Switched PLDS control -
eg_plds_switched_ctrl(.cpp|.py) - tutorial - GLDS fit -
eg_glds_fit.py,test_glds_fit.m - PLDS fit -
eg_plds_fit.py,test_plds_fit.m
RTXI modules:
Latent-switch decoding (HMM) CLOCTools/hmm
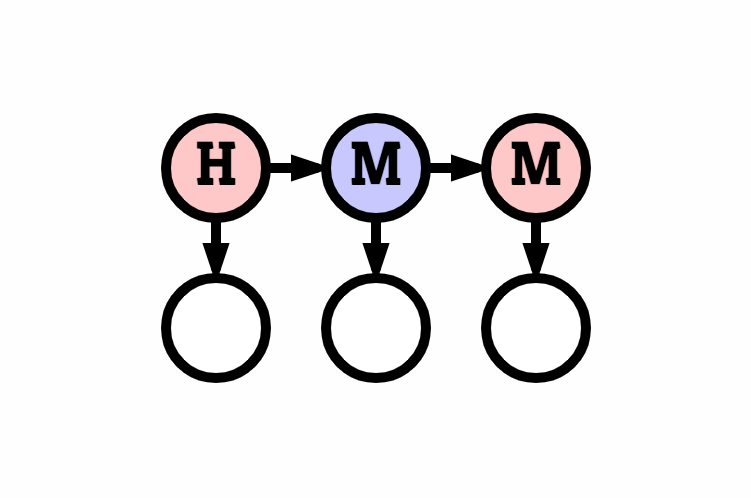
hmm is a simple set of hidden Markov model (HMM) code intended to support control of switching linear dynamical systems. It contains methods for generating and decoding systems with discrete latent states and discrete observed signals.
- code, documentation
- example script with build, simulate, and decode -
test/main.cpp, tutorial - RTXI modules:
State-aware (switched) control of neural dynamics (StAC)
StAC allows for model-based control and estimation of a switched (PLDS) system. It relies on the CLOCTools/ldsCtrlEst and CLOCTools/hmm libraries.
Building a switched control loop is demonstrated at eg_plds_switched_ctrl.cpp, an RTXI wrapper for switched control of a PLDS system suitable for custom decoding is available at rtxi-pldsSwitchedController, and an RTXI wrapper for switched control of a PLDS using an HMM for decoding is available at rtxi-StAC. See also section Satellite:StAC for analysis code and manuscript figures.
Linear quadratic model-predictive control CLOCTools/lqmpc
Model-predictive control (MPC) is a more flexible form of optimal control that allows for constraints and time-varying cost functions. This Python package provides an easy-to-use implementation for the case of linear dynamics and quadratic cost functions, yielding a convex optimization problem which we solve with OSQP.
Implementation tools
Cross-language utilities
CLOCTools makes use of a variety of programming languages. This section lists tools to interface across these languages.
- eg-cpp-library - An example repository for C++ library development
- lds-ctrl-est-pybind - python bindings to ldsCtrlEst core C++ functionality
- lds-ctrl-est-matlab - MATLAB implementations, bindings to ldsCtrlEst
- cpp-matlab-demos - using mex to verify algos in c++ (currently a part of the HMM repository)
Real-time utilities
In order to make use of real time utilities, a variety of wrappers and signal processors are needed. Repositories for these wrappers and signal processors relevant to particular tools are listed in the sections with those tools, while more general repositories for real time utilities are listed in this section.
RTXI, the Real-Time eXperiment Interface, is a hard real-time data acquisition and control application for biological research. At its core is a Real-Time Operating System (RTOS), which uses a modified Linux kernel to provide deterministic control in a variety of experimental settings. RTXI is a fast, free, and open-source system currently used in labs all over the world to probe diverse, technologically challenging problems, such as dynamic probing of ion-channel function, control of cardiac arrhythmia dynamics, and control of deep-brain stimulation patterns. We have repositories with useful tools for interfacing with and configuring RTXI.
- rtxi-settings - RTXI experiment settings
- stimLoader - Stimulus and parameter I/O
RTXI signal processing modules
Tucker Davis Technologies (TDT) have a variety of hardware and software solutions for neuroscience. We have repositories with useful tools for interfacing with TDT hardware and software.
TDT utilities:
- rtxi-tdtSpikes - retrieve spike counts from TDT
- rtxi-tdtLFP - receive local field potential (LFP)
- tdtUDP - A repository for receiving/sending data through Tucker Davis Technologies UDP interface
Related publications
CLOC Tools: A Library of Tools for Closed-Loop Neuroscience
A.A. Willats, M.F. Bolus, K.A. Johnsen, G.B. Stanley, and C.J. Rozell. In prep, 2022.
State-Aware Control of Switching Neural Dynamics
A.A. Willats, M.F. Bolus, C.J. Whitmire, G.B. Stanley, and C.J. Rozell. In prep, 2022.
Closed-Loop Identifiability in Neural Circuits
A. Willats, M. O’Shaughnessy, and C. Rozell. In prep, 2022.
State-space optimal feedback control of optogenetically driven neural activity
M.F. Bolus, A.A. Willats, C.J. Rozell and G.B. Stanley. Journal of Neural Engineering, 18(3), pp. 036006, March 2021.
Design strategies for dynamic closed-loop optogenetic neurocontrol in vivo
M.F. Bolus, A.A. Willats, C.J. Whitmire, C.J. Rozell and G.B. Stanley. Journal of Neural Engineering, 15(2), pp. 026011, January 2018.
Satellite repositories
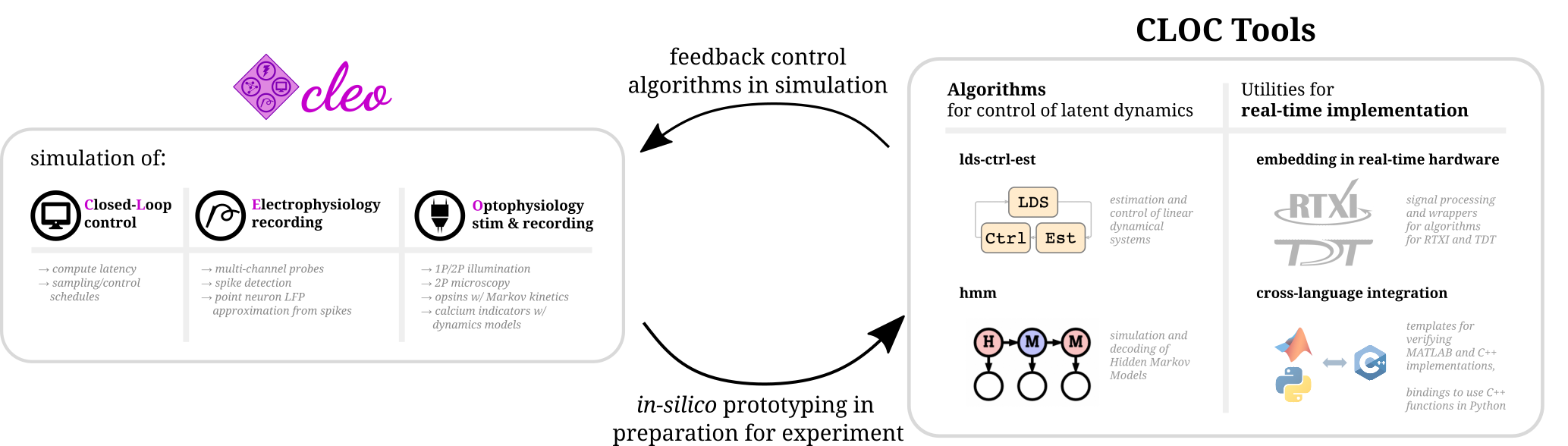
These projects contain related work that doesn’t necessarily fit into the primary focus of CLOCTools or have their own goals separate from CLOCTools. They are listed here for completeness and relevance. Of particular importance is CLEOSim which can serve to test the algorithms in ldsCtrlEst and hmm and to prototype experiments in-silico.
StAC (analysis & manuscript figures)
StAC - State-aware control of switching neural dynamics. StAC applies control, decoding, and estimation to switching models of neural responses. Hidden Markov models and Poisson linear dynamical systems are used to design and characterize feedback in simulation and are compared to non-switching “state-naive” approaches. Real-time implementation is provided by CLOCTools/rtxi-StAC, however CLOCTools/state-aware-control-manuscript contains additional simulations, analysis, and plotting (in MATLAB) for the associated manuscript (in preparation).
Cleo (Closed-Loop, Electrophysiology, and Optophysiology experiment simulation testbed) is a Python package built on the Brian 2 spiking neural network simulator developed bridging theory and experiment for mesoscale neuroscience, facilitating electrode recording, optogenetic stimulation, two-photon imaging, and closed-loop experiments. Cleo can serve a tool for both prototyping experiments and, by simulating perturbation and measurement technolgoy, for producing simulated results that are more comparable to experimental results.
CLINC
CLINC - Closed-loop identifiability in neural circuits. This project proposes a framework for understanding the impact of open and closed-loop interventions in identifying neural circuits. This manuscript (in preparation) summarizes the role of stimulation in circuit inference and demonstrates how to design interventions through simple gaussian network simulations. See also https://github.com/awillats/brian_delayed_gaussian for simulating networks of Gaussian nodes with delayed connections in the Brian2 simulator framework.


